Understanding the Distribution of Mutations along Genomes
Dr. Peter Arndt
Keywords: population genetics, coalescent theory, genome evolution
The genetic information stored in DNA is subject to a variety of mutagenic processes. These processes can change a single base pair resulting in a single nucleotide polymorphism (SNP), insert or delete a stretch of sequence (indels), or result in more complex changes such as inversions or ectopic rearrangements. It is important to have a good understanding of the particular distribution of the effects of all these processes along the DNA. Local patterns will play a role not only in evolution, but also in the development of diseases, such as cancer.
Recent advances in efforts to sequence the whole genome of many individuals in a population and the availability of these data give us the opportunity to reevaluate and extend models of DNA mutation. Our modeling will also incorporate the demographic history of populations based on coalescent theory. This will allow us to resolve complex patterns of DNA change along chromosomes.
Successful candidates will develop projects to study sequencing data from human (or other species) population samples as well as sequencing data from cancer samples. A strong background in dynamical modeling, population genetics, coalescent theory, statistical analysis, computational biology, and machine learning is required. Coding experience in Julia is a bonus.
For more information visit the website of the Evolutionary Genomics Group.
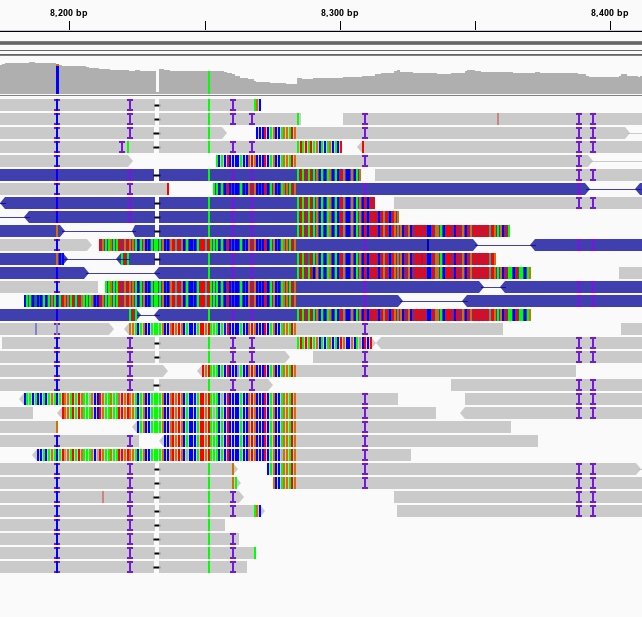
Sequencing reads aligned to the human reference genome as seen in the Integrative Genomics Viewer (IGV). Missmatches and Indels are highlighted using different colors.
