The role of RNA Helicase DHX9 and RNA editase ADAR in RNA processing in neurons
We recently showed that DHX9 is responsible for resolving long double-stranded RNA (dsRNA) that our cells naturally produce during transcription of genes that contain copious amounts of Alu insertions in their intronic regions (Aktas et al. Nature 2017).
Under normal conditions, these sequences are retained in the nucleus and are not allowed to reach the cytoplasm where they can trigger the dsRNA-response and lead to cell death. We further discovered that DHX9, a nuclear enzyme, interacts specifically with the interferon-inducible isoform of ADAR, an enzyme that typically engages viral dsRNA in the cytoplasm. Our results suggest a system of complex checks and balances, whereby our cells tolerate large amounts of nuclear dsRNA through the collective action of DHX9 and ADAR, but still retain the ability to respond to viral assaults in the cytoplasm.
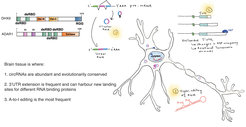
There are several important implications of these results. From an evolutionary perspective, our results suggest that our cells have found ways to tolerate more and more transposable element insertions, as their harmful effects are neutralized by RNA-binding proteins like DHX9 and ADAR, opening the way to even more insertions over time.
In addition to the evolutionary implications, especially with regards to co-evolution of transposons/viruses with the host factors and the ultimate impact of this to the evolution of new traits and speciation through domestication of transposons/viruses, our results point to more practical issues as well: What are the consequences of orderly (e.g. cell division) and disorderly (e.g. aging, physical damage, genetic mutations) breakdown of nuclear envelope that normally shields low-complexity, transposon-riddled intronic RNA that is prone to aggregate formation from the cytoplasm? Does the tolerance to the expansion of transposable elements in our genome come with a hidden cost, which reveals itself as our cells age? What are the roles of enzymes like DHX9 and ADAR in these events?
Our goal is to tackle these questions by first focusing on RNA-binding proteins that play a role in the taming of repetitive elements, such as DHX9, ADAR and hnRNPC. We plan to expand this list, by specifically looking for factors that interact with transposable elements in vivo, and study their broader effect on post-transcriptional regulation of gene expression. Initial approaches will include investigation of fast-cycling cells such as mouse embryonic stem cells (mESc) and terminally differentiated cells such as neurons which will be further expanded into mouse models using tissue specific/conditional knock-out alleles.
References:
DHX9 suppresses RNA processing defects originating from the Alu invasion of human genome Aktas T.*; Ilik I.A.*; Maticzka D.; Bhardwaj V.; Rodrigues C.P.; Manke T.; Backofen R.; Akhtar A.; Nature,544 (7648), 115-119, 2017
*equal contribution
