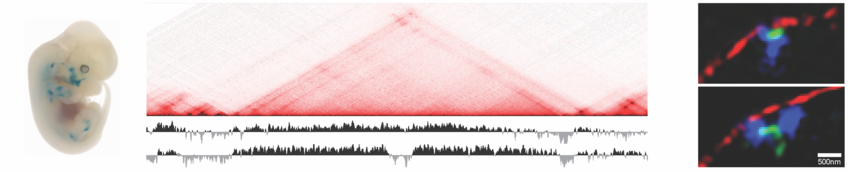
Gene Regulation from the Nuclear Envelope
Many developmental genes acquire intricate patterns of expression by physically contacting and communicating with enhancers located within conserved regulatory landscapes. To facilitate this communication, regulatory landscapes are highly structured in 3D space and must transition between fundamentally different states of repression and activation during development. However, such transformations are complex with landscapes seemingly altering transcription, chromatin state, structure and attachment to the repressive nuclear envelope near simultaneously. Consequently, the significant problem remains; which alteration directs loci to reconfigure and does this itself control gene expression?
New technological innovations now provide an unprecedented opportunity to address these questions. Low input structure and chromatin mapping techniques allow landscape behaviors to be contrasted across evolutionary time in non-standard model organisms. Super-resolution OligoPAINT FISH microscopy and combined single-cell sequencing methods allow the transcription, structure, and nuclear envelope-attachment of such regulatory landscapes to then be directly measured at individual alleles. Finally, complimentary CRISPR-genome engineering and rapid mouse-generation approaches enable examined landscapes to be genetically disrupted or transplanted and then functionally examined during development in vivo.
We now integrate and further refine these approaches to interrogate and disrupt regulatory landscapes and their transitions during in vivo mouse limb and retina development. In this way, we hope to understand how single loci are regulated in time and space and how this is sculpted to alter development in evolution and disease.